Flex Funds Project: metaTraits
A large-scale integration of microbial phenotypic trait information (metaTraits)
Project Start and End Date
2023-03-01 - 2024-02-29
Short project summary
Microbial phenotypic traits offer vital insights into the roles microbes play in natural ecosystems and human health. However, the fragmentation and dispersion of this information across various public databases leave the vast potential of these invaluable culture collections largely unexploited, restricting their application in annotating metagenomic datasets and derived genomes. Furthermore, because traditional databases rely heavily on cultivated isolates, there is a significant data gap for the vast majority of microbial taxa that remain uncultured and are only represented by metagenome-assembled genomes (MAGs).
To overcome this, the metaTraits project aims to conduct a large-scale integration of these public repositories into a single, comprehensive resource. Our primary objective is to make microbial phenotypic trait information highly accessible, interoperable, and reusable for the broader scientific community.
The project aims:
- To develop an intuitive web service that facilitates the user-friendly search, exploration, and visualization of microbial traits, complete with transparent data provenance.
- To enhance the NFDI4Microbiota platform’s analytical capabilities by engineering automated workflows that empower researchers, regardless of their technical expertise, to seamlessly annotate their own genomic data and taxonomic profiles with rich phenotypic insights.
Graphical abstract
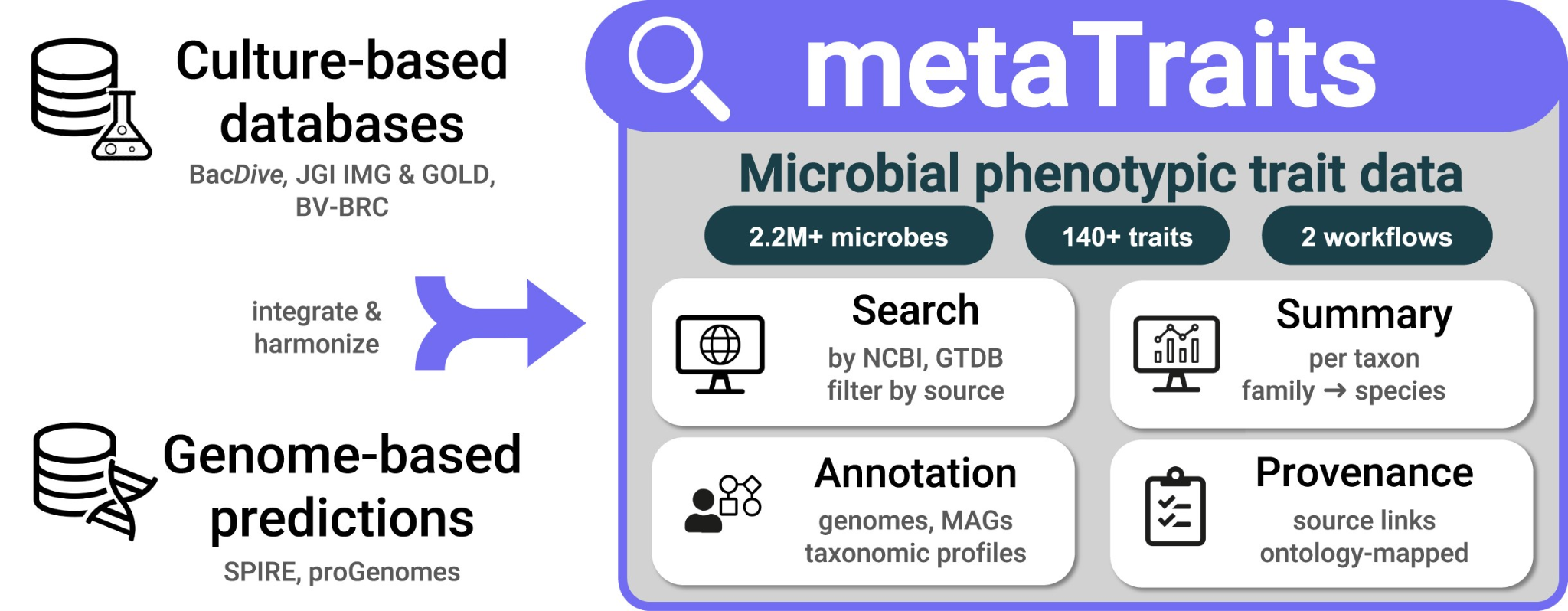
Brief summary of the main results and conclusion
The metaTraits project successfully achieved its goal of integrating fragmented microbial phenotypic trait data into a unified, accessible, and FAIR-compliant resource. We developed a robust framework that harmonizes culture-based trait data from diverse public repositories and augments it with sequence-based trait predictions for both isolate genomes and metagenome-assembled genomes (MAGs), which effectively bridges the critical data gap for uncultured taxa. metaTraits features a dedicated website at metaTraits.embl.de offering user-friendly search and exploration tools. The service aggregates isolate-level trait data across databases to generate trait estimates at the taxonomic species level and above, displaying these estimates and their distributions with relevant context. Additionally, metaTraits introduces two automated workflows via its web interface for annotating user-submitted microbiome data with phenotypic trait estimates. One workflow is tailored for annotating microbial genomes (both isolates and MAGs), and a second workflow allows for bulk annotating entire microbial populations based on taxonomic profiles from tools such as mOTUS or MetaPhlAn. By centralizing this data, metaTraits can therefore not only serve as a quick reference tool for researchers but can also be seamlessly integrated with software widely used by the microbiome research community.
Achievements
- Publication of the metaTraits webservice: “Podlesny, D., Kim, C. Y., Robbani, S. M., Schudoma, C., Fullam, A., Reimer, L. C., Koblitz, J., Schober, I., Iyappan, A., Van Rossum, T., Schiller, J., Grekova, A., Kuhn, M., & Bork, P. (2026). metaTraits: a large-scale integration of microbial phenotypic trait information. Nucleic acids research, 54(D1), D835–D841. https://doi.org/10.1093/nar/gkaf1241”
- metaTraits web service live at metaTraits.embl.de
- porTraits workflow for the annotation of (metagenome-) assembled genomes with microbial phenotypic trait predictions. Live on metaTraits.embl.de, CloWM and GitHub
- sumTraits workflow for the bulk annotation of taxonomic profiles and the calculation of community-weighted trait averages. Live on metaTraits.embl.de
- metaTraits service enabled a high-impact publication of a planetary-scale microbiome analysis: “Kim, C. Y., Podlesny, D., Schiller, J., Khedkar, S., Fullam, A., Orakov, A., Schudoma, C., Robbani, S. M., Grekova, A., Kuhn, M., & Bork, P. (2026). Planetary microbiome structure and generalist-driven gene flow across disparate habitats. Cell, S0092-8674(25)01500-4. Advance online publication. https://doi.org/10.1016/j.cell.2025.12.051”
- Featured as an NFDI success story https://www.nfdi.de/successstories/
Project Members
Dr. Daniel Podlesny
ORCID ID: 0000-0002-5685-0915
European Molecular Biology Laboratory (EMBL)
Project Lead
Mahdi Robbani
ORCID ID: 0000-0003-0161-0559
European Molecular Biology Laboratory (EMBL)
Dr. Christian Schudoma
ORCID ID: 0000-0003-1157-1354
European Molecular Biology Laboratory (EMBL)
Dr. Thea Van Rossum
ORCID ID: 0000-0002-3598-5001
European Molecular Biology Laboratory (EMBL)
Dr. Lorenz Reimer
ORCID ID: 0000-0002-7805-0660
German Collection of Microorganisms and Cell Cultures (DSMZ)
Dr. Isabel Schober
ORCID ID: 0000-0002-4894-1913
German Collection of Microorganisms and Cell Cultures (DSMZ)
Dr. Julia Koblitz
ORCID ID: 0000-0002-7260-2129
German Collection of Microorganisms and Cell Cultures (DSMZ)
Keywords
phenotypic traits
integration
metagenomics
microbiome