Flex Funds Project: EnterArchaeo
Enhancing training, resource development, and analytical reproducibility in ancient microbiome research (EnterArchaeo)
Project Start and End Date
2025-01-01 - 2025-12-31
Short project summary
Ancient DNA has emerged as a powerful tool for revealing the hidden and often surprising evolutionary histories of microbiomes. Recent advances in the application of metagenomics to ancient DNA data is now allowing the recovery of extinct and novel microbes and their genes. The level of ancient microbiome data being produced promises to establish an astonishing record of the microbial genetic past, but this requires investment in training, resource development, and analytical reproducibility. This project proposed three pillars to address these necessities to maximise the value of ancient microbiome data:
- Implementation of and training in the development and usage of high-throughput reproducible Nextflow metagenomic pipelines, as well as analysis of ancient microbiome data
- Support development of dedicated gold-standard metadata reporting schemas for ancient DNA for improved data accessibility and findability.
- Contribute to the continued maintenance and extensions of existing and new open-source nf-core pipelines for both modern and ancient microbiome research. All pillars are designed to widely disseminate knowledge and upskill researchers following FAIR data and reproducible data analysis practices to ensure maximum value and uptake of (ancient) microbiome data.
Graphical abstract
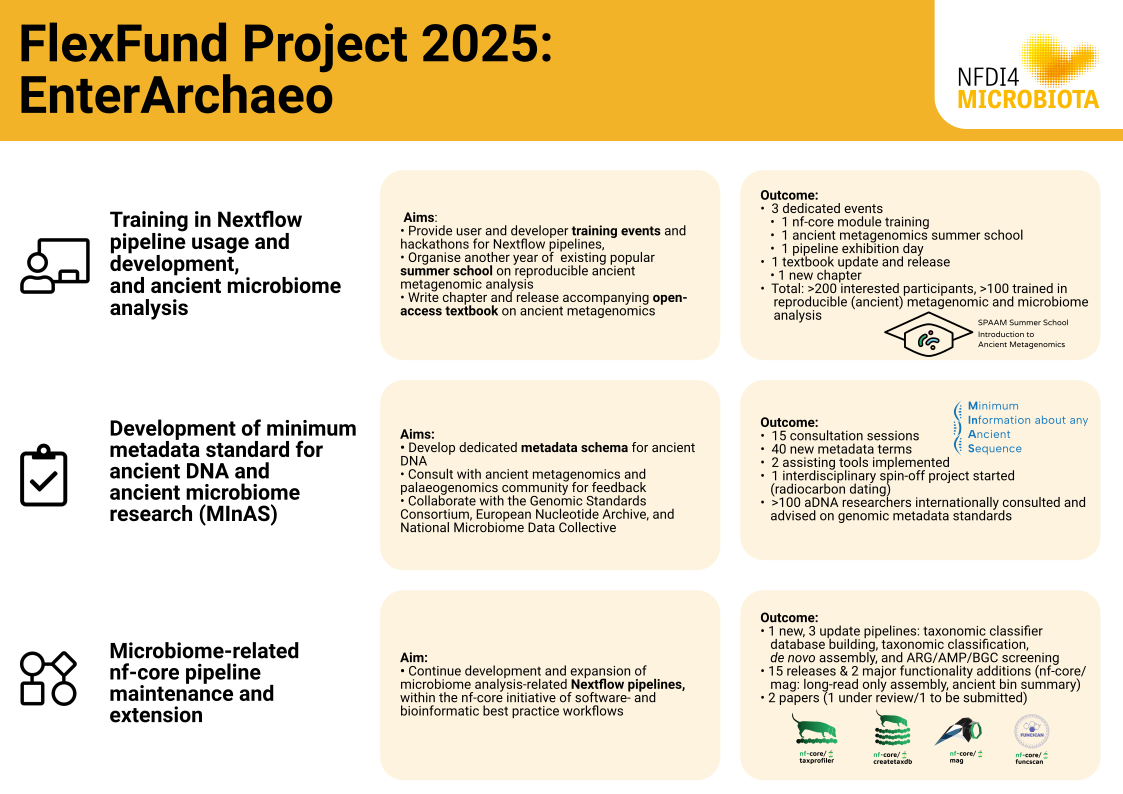
Brief summary of the main results and conclusion
The project saw a total of three dedicated events for training in writing Nextflow code, one bioinformatic summer school on ancient metagenomics, and one community exhibition day for Nextflow meta-*omics pipelines. Across the three dedicated events, there were >200 applicants, and a total >100 trained on the different topics. One new textbook chapter on NGS sequencing of ancient metagenomic samples was completed, and included in the also released latest version of the summer school companion online and open-access textbook. 15 consultation workshops were carried out with >100 globally distributed ancient DNA researchers as well as with members of the Genomic Standard Consortium, European Nucleotide Archive, and National Microbiome Data Collaborative Data. In total 40 new ancient DNA specific metadata terms were developed and refined for the MInAS metadata standard, to be merged with the existing MIxS standard mid-2026. Two helper tools were developed, and one interdisciplinary spin-off project on radiocarbon-dating metadata reporting was initiated. Four nf-core best-practice Nextflow pipelines were developed during the project - one new pipeline for taxonomic classifier database building had its first release. Three pipelines for taxonomic classification, metagenomic de novo assembly, and for antimicrobial gene, peptide, and biosynthetic gene cluster screening pipelines were maintained. This included bug fixes, version updates, and for nf-core/mag, major new functionality (long-read only assembly, and improved ancient DNA authentication summary). Two academic papers were written for nf-core/mag, one under review and the other to be submitted.
Overall the project disseminated open-science and FAIR data best-practices to interdisciplinary audiences within Germany and internationally, and saw the sustained development of popular high-quality microbiome related tooling and FAIR data resources.
<< Back to all past Flex FundsProject Members
Dr. James Fellows Yates
ORCID ID: 0000-0001-5585-6277
Leibniz Institute for Natural Product Research and Infection Biology - Hans Knöll Institute (Leibniz-HKI)
Prof. Christina Warinner
ORCID ID: 0000-0002-4528-5877
Leibniz Institute for Natural Product Research and Infection Biology - Hans Knöll Institute (Leibniz-HKI)
Keywords
ancient microbiome
training
resource development
reproducibility