Flex Funds Project: catch-M-all
Metadata capture from data generation to submission (catch-M-all)
Project Start and End Date
2026-01-01 - 2026-12-31
Short project summary
Millions of microbe-related omics datasets are submitted to public repositories such as the European Nucleotide Archive (ENA), where high-quality metadata is essential to enable reuse and accelerate microbiological research and innovation. However, a major bottleneck is the attrition of metadata between sample collection, sequencing, and final submission. Complex submission workflows and the perceived burden of proper curation often discourage researchers, resulting in records with incomplete, inconsistent, or missing biological metadata.
While metadata is frequently treated as an administrative afterthought, this use case promotes that high-quality, standardized metadata is captured and curated from the very start of the data generation process. By connecting existing tools developed within the NFDI4Microbiota consortium, we will conceptualize a robust, generalized framework that ensures metadata is collected, FAIR-compliant, actively curated (with the support of data stewards), and securely linked to sequencing data right upon requesting data from sequencing providers.
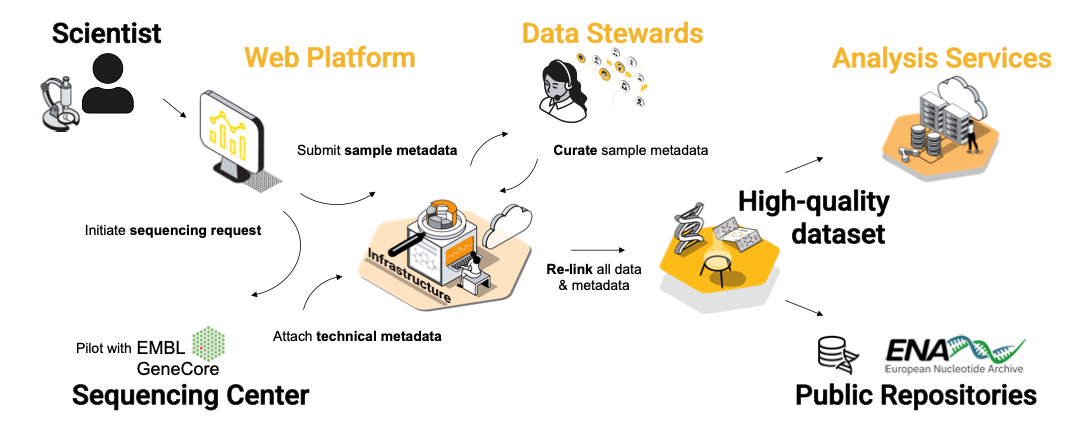
To allow for this end-to-end solution, a web platform that enables the initiation of sequencing requests with cooperating sequencing centers, will also capture and validate biological metadata upfront, assigning unique identifiers to each sample. To minimize data protection concerns and administrative overhead, only these identifiers and minimal technical instructions are passed to sequencing centers (e.g., EMBL GeneCore) for the initiation of data generation. In parallel, NFDI4Microbiota data stewards curate and enrich the biological metadata. Once sequencing is complete, raw data and technical metadata are returned and automatically re-linked to the curated biological profiles via the shared identifiers. The result is a fully annotated dataset with high-quality metadata, ready for downstream analysis (e.g., through CloWM) and submission to public repositories.
To validate this framework, the EMBL GeneCore sequencing facility will serve as a real-world testing ground, demonstrating the feasibility of our cohesive data brokerage pipeline in a high-throughput environment.
Graphical abstract
How you can contribute
You are a …
- microbiologist planning to sequence with EMBL GeneCore: Get in touch with our development team to test the data workflow and upfront metadata capture..
Planned output
Integrated data brokerage pipeline
- To conceptualize the connection of existing NFDI4Microbiota tools (such as ARUNA, CloWM, Broker4Microbiota, ENA wizard tool) into a cohesive data brokerage pipeline.
- A decoupled workflow that successfully separates biological metadata collection from technical metadata generation, enabling the curation of biological metadata while sequencing is still in progress.
- An automated mechanism for re-linking raw sequencing data and technical metadata to curated biological metadata using unique upfront identifiers.
- A collaborative implementation of this framework with NFDI4Microbiota partners.
Real-world pilot datasets
- A piloted framework tested at the EMBL GeneCore facility to resolve friction for both users and sequencing providers.
- A (pseudo)-generated, fully annotated, and FAIR-compliant dataset ready for ENA submission.
- A finalized, scalable blueprint suitable for adoption and testing by other sequencing centers.
Project Members
Dr. Daniel Podlesny
ORCID ID: 0000-0002-5685-0915
European Molecular Biology Laboratory (EMBL)
Project Lead
Mahdi Robbani
ORCID ID: 0000-0003-0161-0559
European Molecular Biology Laboratory (EMBL)
Team Member
Dr. Christian Schudoma
ORCID ID: 0000-0003-1157-1354
European Molecular Biology Laboratory (EMBL)
Team Member
Dr. Vladimir Benes
ORCID ID: 0000-0002-0352-2547
European Molecular Biology Laboratory (EMBL)
Team Member
Dr. Adrian Forsythe
ORCID ID:
European Molecular Biology Laboratory (EMBL)
Team Member
Keywords
sequencing center
data generation
metadata curation
data broker